Massive parallelism computation by Spark.
Abstract: Text analysis is a very computation-resource-cost activity with high memory requirement, but normal single computer, which with limited CPU and memory, can not handle it fast. With Spark we can do it better, spark is known well for streaming iterative computation and distributed memory building on large scale of clusters. And with the research paper dataset from PubMed, our idea becomes to category documents by calculating betweenness centrality of words from text network with Spark. The intensive comparative evaluation has been made with three typical indexes in network analysis which are degree centrality, closeness centrality, and betweenness centrality. For a documentation, if the topic of documentation is biology, then a lot of compound nouns with the word “biology” should appear frequently. Therefore, the topics of documents can be presumed by extracting the compound nouns from the document and by focusing on nouns that consist them. The measurement is the appearance ratio of category name at top n words with high centrality.
Keywords: Spark, BFS, Shortest Path, Betweenness centrality, Text Analysis
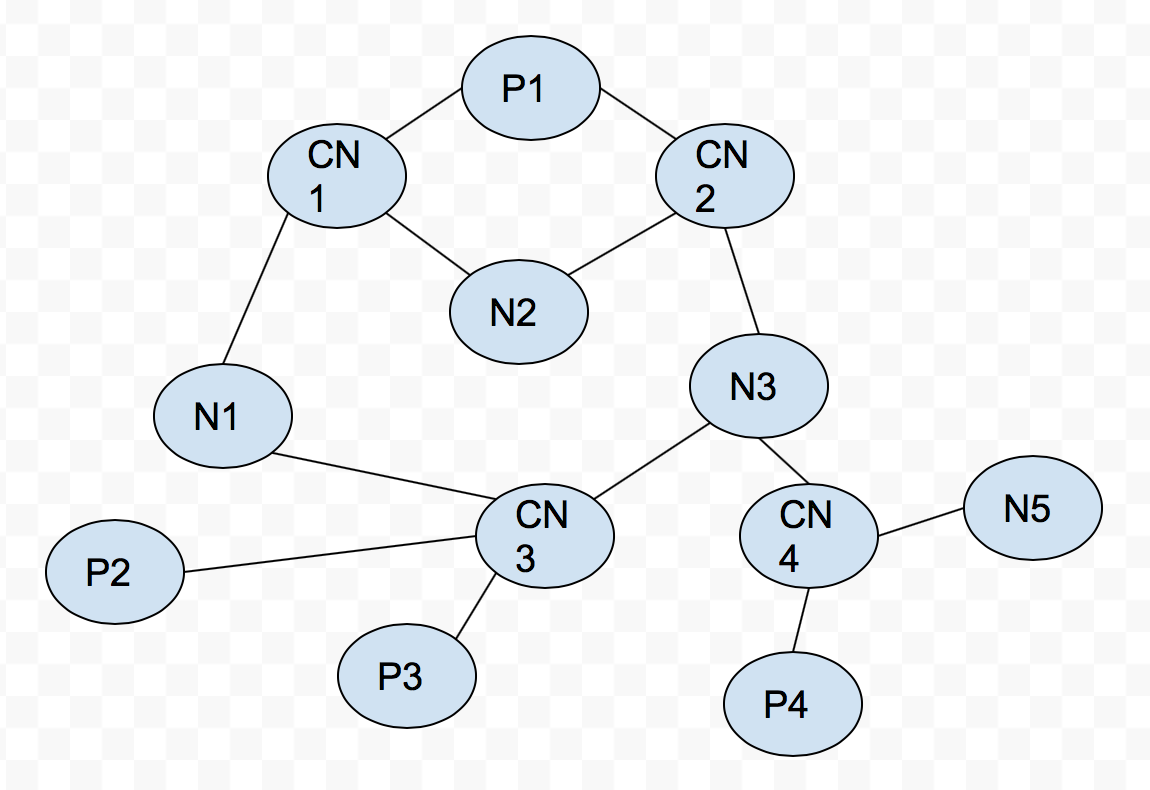
P means paper node, CN is compound nouns node, N is noun node.
class Node {
long id,
Object attr
}
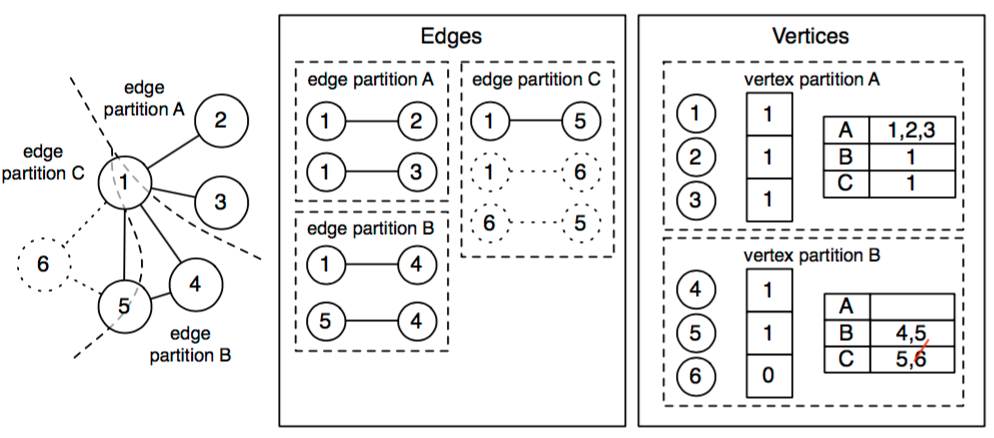
The vertices are partitioned by id. Within each vertex partition, the routing table stores for each edge partition the set of vertices present. Vertex 6 and adjacent edges (shown with dotted lines) have been restricted from the graph, so they are removed from the edges and the routing table. Vertex 6 remains in the vertex partitions, but it is hidden by the bitmask.
Betweenness centrality is an indicator of a node's centrality in a network. It is equal to the number of shortest paths from all vertices to all others that pass through that node. A node with high betweenness centrality has a large influence on the transfer of items through the network, under the assumption that item transfer follows the shortest paths.
g(v) = ∑ s ≠ v ≠ t σ s,t (v) σ s,twhere σ s,t is the number of shortest (s, t)-paths, and σ s,t (v) is the number of those paths passing through some node v other than s, t. If s = t, σ s,t = 1, and if v in {s, t}, σ s,t (v) = 0
Always when running in single thread, we use Dijkstra or Bell-Ford algorithm to find shortest pathes, but in paralle situation, the former 2 algorithm is not easy to implement. But we can use parallel Breadth-First Search which maps process on each node to find all shortest pathes.
class MAPPER
method Map(VertexId id, Node N)
d <- N.Distance
EMIT(id, [])
for n in N.Neiborhood do
EMIT(id, list+n)
class REDUCER
method Reduce(VertexId id, Array[] list)
for path in list
if shortest(path)
result <- path
EMIT(id, result)
The idea of algorithm we implemented is from Ulrik Brandes
We test the program on CCR HPC in Buffalo, with computation resources range from 1 to 32. The datesets is from PubMed, we picked 20GB plain file to test.