GeoStatTools provides geostatistical tools for random field generation and variogram estimation based on many readily provided and even user-defined covariance models.
The package can be installed via pip. On Windows you can install WinPython to get Python and pip running.
pip install gstools
You can find the documentation under geostat-framework.readthedocs.io.
The documentation also includes some tutorials, showing the most important use cases of GSTools, which are
Some more examples are provided in the examples folder.
The core of this library is the generation of spatial random fields. These fields are generated using the randomisation method, described by Heße et al. 2014.
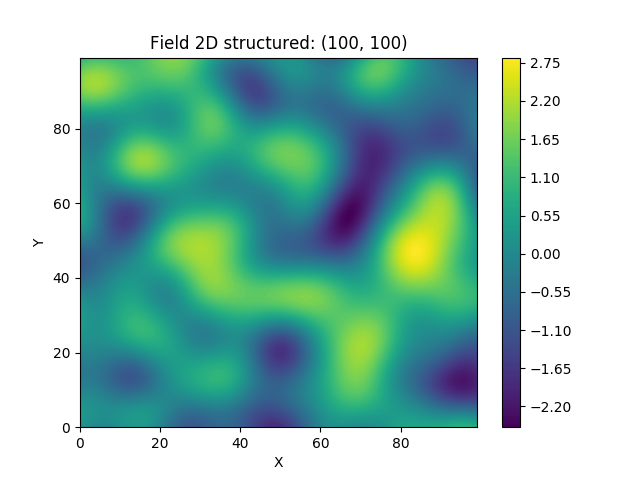
This is an example of how to generate a 2 dimensional spatial random field with a gaussian covariance model.
from gstools import SRF, Gaussian
import matplotlib.pyplot as plt
# structured field with a size 100x100 and a grid-size of 1x1
x = y = range(100)
model = Gaussian(dim=2, var=1, len_scale=10)
srf = SRF(model)
field = srf((x, y), mesh_type='structured')
plt.imshow(field)
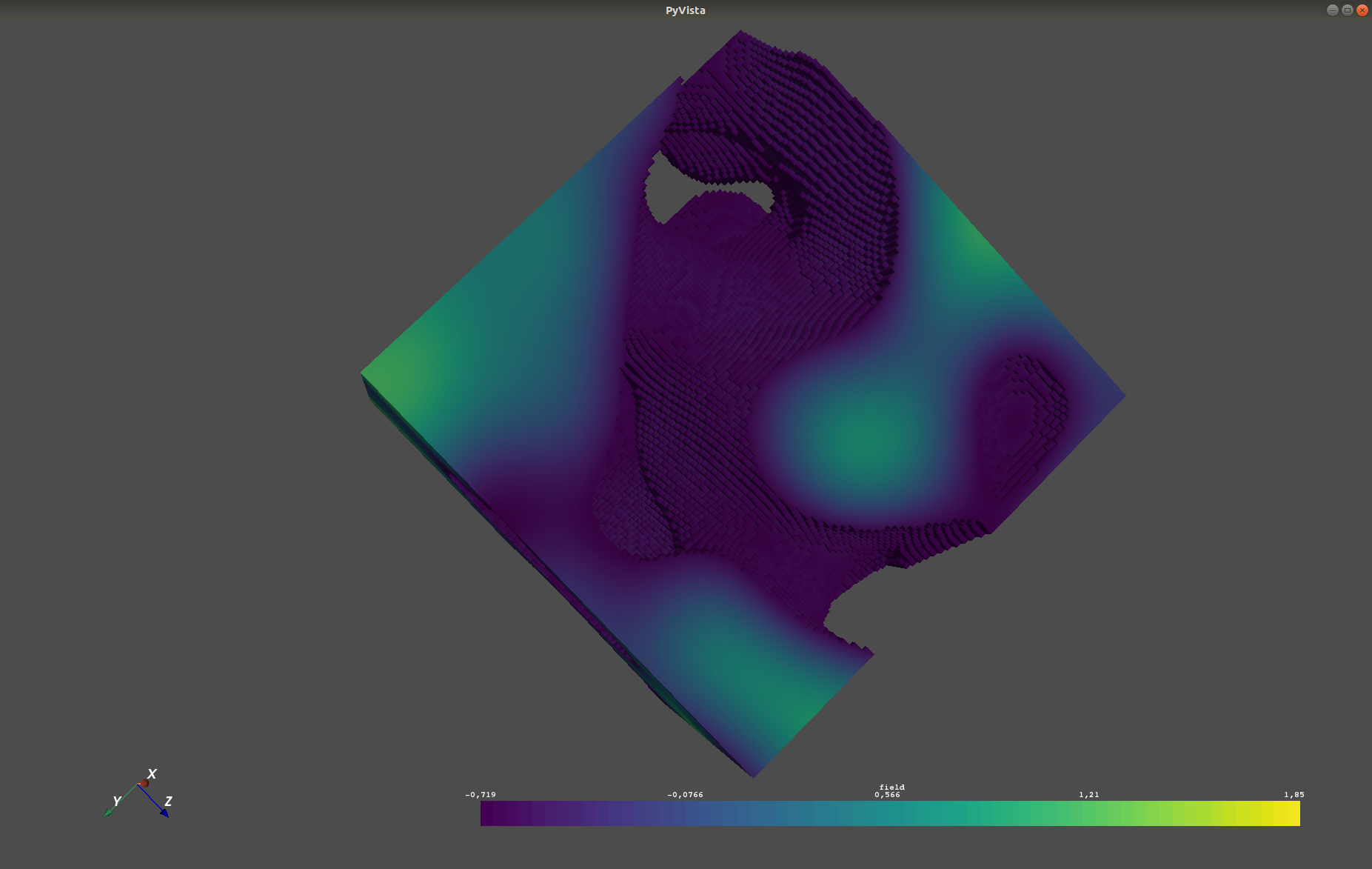
plt.show()A similar example but for a three dimensional field is exported to a VTK file, which can be visualised with ParaView.
from gstools import SRF, Gaussian, vtk_export
import matplotlib.pyplot as pt
# structured field with a size 100x100x100 and a grid-size of 1x1x1
x = y = z = range(100)
model = Gaussian(dim=3, var=0.6, len_scale=20)
srf = SRF(model)
field = srf((x, y, z), mesh_type='structured')
vtk_export('3d_field', (x, y, z), field, mesh_type='structured')GSTools also implements truncated power law variograms, which can be represented as a superposition of scale dependant modes in form of standard variograms, which are truncated by an upper lengthscale lu.
This example shows the truncated power law based on the stable model and is given by
which gives Gaussian modes for alpha=2 or exponential modes for alpha=1
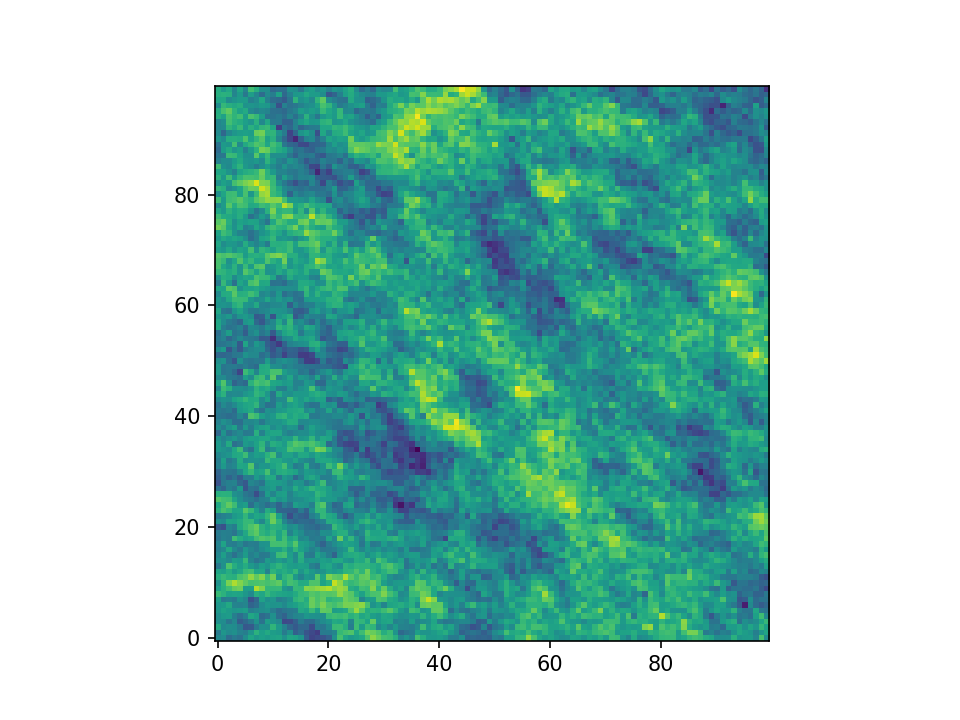
This results in:
import numpy as np
import matplotlib.pyplot as plt
from gstools import SRF, TPLStable
x = y = np.linspace(0, 100, 100)
model = TPLStable(
dim=2, # spatial dimension
var=1, # variance (C is calculated internally, so that the variance is actually 1)
len_low=0, # lower truncation of the power law
len_scale=10, # length scale (a.k.a. range), len_up = len_low + len_scale
nugget=0.1, # nugget
anis=0.5, # anisotropy between main direction and transversal ones
angles=np.pi/4, # rotation angles
alpha=1.5, # shape parameter from the stable model
hurst=0.7, # hurst coefficient from the power law
)
srf = SRF(model, mean=1, mode_no=1000, seed=19970221, verbose=True)
field = srf((x, y), mesh_type='structured', force_moments=True)
# show the field in xy coordinates
plt.imshow(field.T, origin="lower")
plt.show()The spatial structure of a field can be analyzed with the variogram, which contains the same information as the covariance function.
All covariance models can be used to fit given variogram data by a simple interface.
This is an example of how to estimate the variogram of a 2 dimensional unstructured field and estimate the parameters of the covariance model again.
import numpy as np
from gstools import SRF, Exponential, Stable, vario_estimate_unstructured
from gstools.covmodel.plot import plot_variogram
import matplotlib.pyplot as plt
# generate a synthetic field with an exponential model
x = np.random.RandomState(19970221).rand(1000) * 100.
y = np.random.RandomState(20011012).rand(1000) * 100.
model = Exponential(dim=2, var=2, len_scale=8)
srf = SRF(model, mean=0, seed=19970221)
field = srf((x, y))
# estimate the variogram of the field with 40 bins
bins = np.arange(40)
bin_center, gamma = vario_estimate_unstructured((x, y), field, bins)
plt.plot(bin_center, gamma)
# fit the variogram with a stable model. (no nugget fitted)
fit_model = Stable(dim=2)
fit_model.fit_variogram(bin_center, gamma, nugget=False)
plot_variogram(fit_model, x_max=40)
# output
print(fit_model)
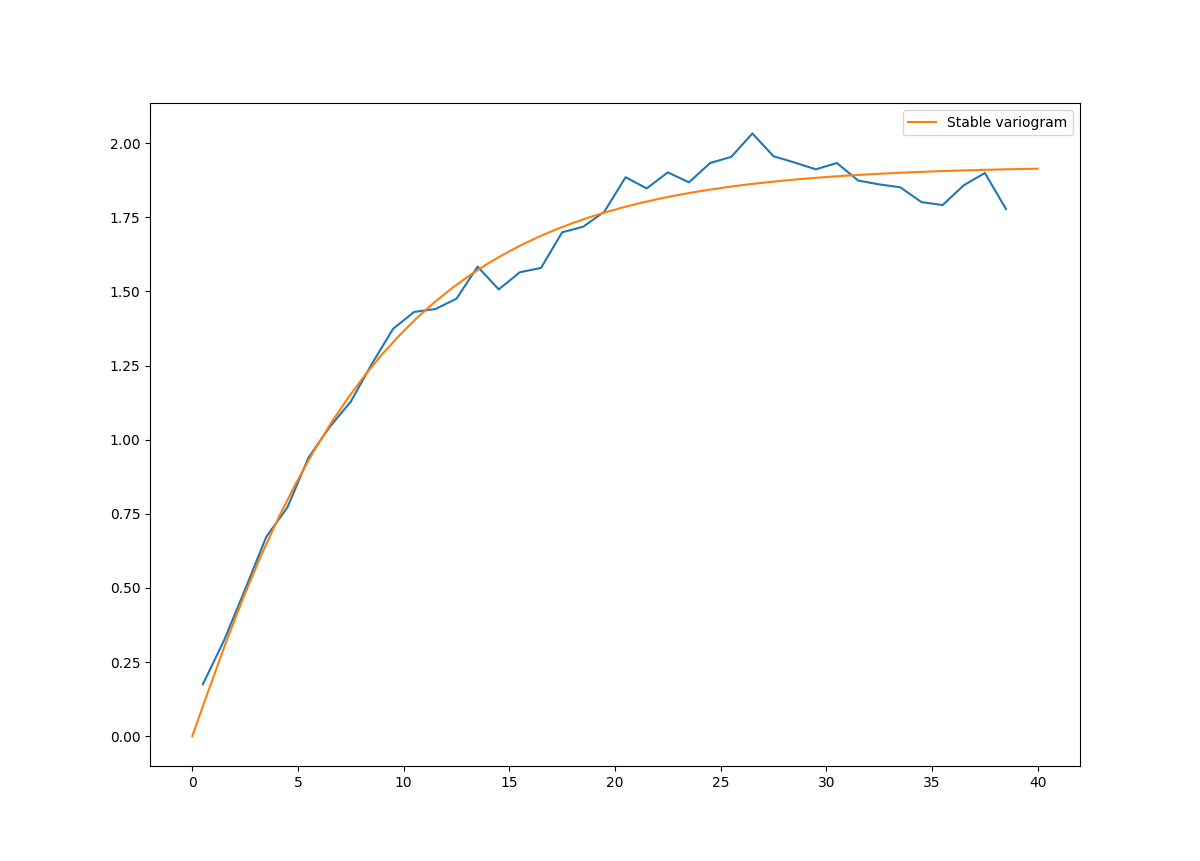
plt.show()Which gives:
Stable(dim=2, var=1.92, len_scale=8.15, nugget=0.0, anis=[1.], angles=[0.], alpha=1.05)One of the core-features of GSTools is the powerfull CovModel class, which allows to easy define covariance models by the user.
Here we reimplement the Gaussian covariance model by defining just the correlation function:
from gstools import CovModel
import numpy as np
# use CovModel as the base-class
class Gau(CovModel):
def correlation(self, r):
return np.exp(-(r/self.len_scale)**2)And that's it! With Gau you now have a fully working covariance model,
which you could use for field generation or variogram fitting as shown above.
Have a look at the documentation for further information on incorporating optional parameters and optimizations.
After you have created a field, you may want to save it to file, so we provide a handy VTK export routine:
from gstools import SRF, Gaussian, vtk_export
x = y = range(100)
model = Gaussian(dim=2, var=1, len_scale=10)
srf = SRF(model)
field = srf((x, y), mesh_type='structured')
vtk_export("field", (x, y), field, mesh_type='structured')Which gives a RectilinearGrid VTK file field.vtr.
You can contact us via info@geostat-framework.org.
GPL © 2018-2019